CatPhan#
Overview#
The CT module automatically analyzes DICOM images of a CatPhan 504, 503, 600, Quart DVT, or ACR phantoms acquired when doing CBCT or CT quality assurance. It can load a folder or zip file that the images are in and automatically correct for translational and rotational errors. It can analyze the HU regions and image scaling (CTP404), the high-contrast line pairs (CTP528) to calculate the modulation transfer function (MTF), the HU uniformity (CTP486), and Low Contrast (CTP515) on the corresponding slices.
For ACR and Quart phantoms, the equivalent sections are analyzed where applicable even though each module does not have an explicit name. Where intuitive similarities between the phantoms exist, the library usage is the same.
Features:
Automatic phantom registration - Your phantom can be tilted, rotated, or translated–pylinac will automatically register the phantom.
Automatic testing of all major modules - Major modules are automatically registered and analyzed.
Any scan protocol - Scan your CatPhan with any protocol; even scan it in a regular CT scanner. Any field size or field extent is allowed.
Running the Demo#
To run one of the CatPhan demos, create a script or start an interpreter and input:
from pylinac import CatPhan504
cbct = CatPhan504.run_demo() # the demo is a Varian high quality head scan
(Source code, png, hires.png, pdf)
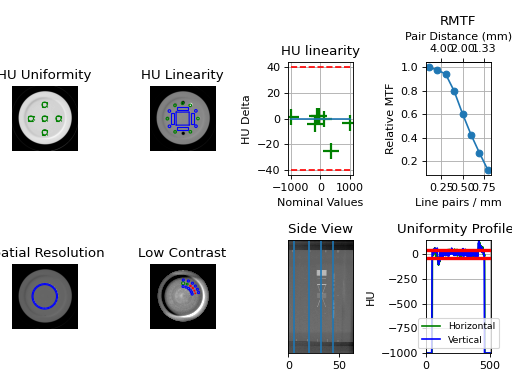
Results will be also be printed to the console:
- CatPhan 504 QA Test -
HU Linearity ROIs: Air: -998.0, PMP: -200.0, LDPE: -102.0, Poly: -45.0, Acrylic: 115.0, Delrin: 340.0, Teflon: 997.0
HU Passed?: True
Low contrast visibility: 3.46
Geometric Line Average (mm): 49.95
Geometry Passed?: True
Measured Slice Thickness (mm): 2.499
Slice Thickness Passed? True
Uniformity ROIs: Top: 6.0, Right: -1.0, Bottom: 5.0, Left: 10.0, Center: 14.0
Uniformity index: -1.479
Integral non-uniformity: 0.0075
Uniformity Passed?: True
MTF 50% (lp/mm): 0.56
Low contrast ROIs "seen": 3
As well, you can plot and save individual pieces of the analysis such as linearity:
(Source code, png, hires.png, pdf)
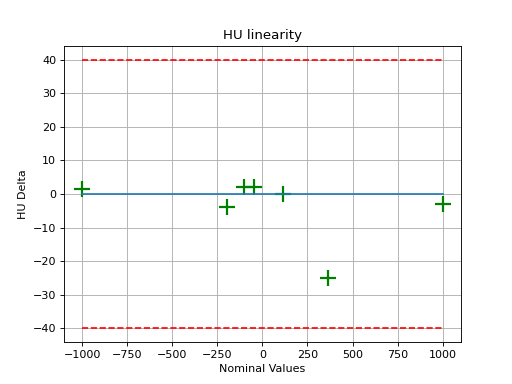
Or the rMTF:
cbct.plot_analyzed_subimage("rmtf")
(Source code, png, hires.png, pdf)
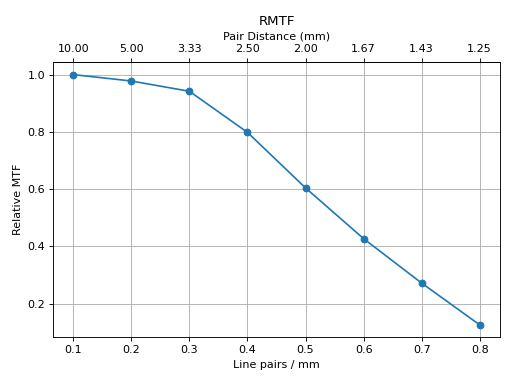
Or generate a PDF report:
cbct.publish_pdf("mycbct.pdf")
Image Acquisition#
Acquiring a scan of a CatPhan has a few simple requirements:
The field of view must be larger than the phantom diameter + a few cm for clearance.
The phantom should not be touching any edge of the FOV.
The phantom shouldn’t touch the couch or other high-HU objects. This may cause localization issues finding the phantom. If the phantom doesn’t have an associated cradle, setting it on foam or something similar is recommended.
All modules must be visible.
Warning
This can cause strange results if not all modules are scanned. Furthermore, aligning axially to the white dots on the side of the catphan will not catch the inferior modules on a typical CBCT scan. We suggest aligning to the center of the HU module (the module inferior to the white dots) when acquiring via CBCT.
Typical Use#
CatPhan analysis as done by this module closely follows what is specified in the CatPhan manuals, replacing the need for manual measurements.
There are 4 CatPhan models that pylinac can analyze: CatPhan504, CatPhan503, & CatPhan600, &
CatPhan604, each with their own class in
pylinac. Let’s assume you have the CatPhan504 for this example. Using the other models/classes is exactly
the same except the class name.
from pylinac import CatPhan504 # or import the CatPhan503 or CatPhan600
The minimum needed to get going is to:
Load images – Loading the DICOM images into your CatPhan object is done by passing the images in during construction. The most direct way is to pass in the directory where the images are:
cbct_folder = r"C:/QA Folder/CBCT/June monthly" mycbct = CatPhan504(cbct_folder)
or load a zip file of the images:
zip_file = r"C:/QA Folder/CBCT/June monthly.zip" mycbct = CatPhan504.from_zip(zip_file)
You can also use the demo images provided:
mycbct = CatPhan504.from_demo_images()
Analyze the images – Once the folder/images are loaded, tell pylinac to start analyzing the images. See the Algorithm section for details and
analyze`()for analysis options:mycbct.analyze()
View the results – The CatPhan module can print out the summary of results to the console as well as draw a matplotlib image to show where the samples were taken and their values:
# print results to the console print(mycbct.results()) # view analyzed images mycbct.plot_analyzed_image() # save the image mycbct.save_analyzed_image("mycatphan504.png") # generate PDF mycbct.publish_pdf( "mycatphan.pdf", open_file=True ) # open the PDF after saving as well.
Custom HU values#
New in version 3.16.
By default, expected HU values are drawn from the values stated in the manual.
It’s possible to override one or more of the HU values of the CatPhan modules however. This is useful if you have a custom CatPhan or
know the exact HU values of your phantom. To do so, pass in a dictionary of the HU values to the expected_hu_values parameter.
The keys should be the name of the material and the values should be the HU value.
from pylinac import CatPhan504
# overrides the HU values of the Air and PMP regions
mycbct = CatPhan504(...)
mycbct.analyze(..., expected_hu_values={"Air": -999, "PMP": -203})
mycbct.plot_analyzed_image()
Note
Not all materials need to be overridden. Only the ones you want to override need to be passed in.
Keys#
The keys to override for CatPhan504 and CatPhan503 are listed below along with the default value:
Air: -1000PMP: -196LDPE: -104Poly: -47Acrylic: 115Delrin: 365Teflon: 1000
The CatPhan600 has the above keys as well as:
Vial: 0
The CatPhan604 has the original keys as well as:
50% Bone: 72520% Bone: 237
Slice Thickness#
New in version 3.12.
When measuring slice thickness in pylinac, slices are sometimes combined depending on the slice thickness. Scans with thin slices and low mAs can have very noisy wire ramp measurements. To compensate for this, pylinac will average over 3 slices (+/-1 from CTP404) if the slice thickness is <3.5mm. This will generally improve the statistics of the measurement. This is the only part of the algorithm that may use more than one slice.
If you’d like to override this, you can do so by setting the padding (aka straddle) explicitly.
The straddle is the number of extra slices on each side of the HU module slice to use for slice thickness determination.
The default is auto; set to an integer to explicitly use a certain amount of straddle slices. Typical
values are 0, 1, and 2. So, a value of 1 averages over 3 slices, 2 => 5 slices, 3 => 7 slices, etc.
Note
This technique can be especially useful when your slices overlap.
from pylinac import CatPhan504 # applies to all catphans
ct = CatPhan504(...)
ct.analyze(..., thickness_slice_straddle=0)
...
Noise Power Spectrum#
New in version 3.19.
The noise power spectrum (NPS) is a measure of the noise in the image using FFTs. It was added to comply with French regulations. It is calculated on the uniformity module (CTP486). Pylinac will provide the most populous frequency and the average power of the NPS.
from pylinac import CatPhan504
ct = CatPhan504(...)
ct.analyze(...)
ct.ctp486.avg_noise_power
ct.ctp486.max_noise_power_frequency
# plot the NPS
ct.ctp486.plot_noise_power_spectrum()
The resulting plot will look like so:
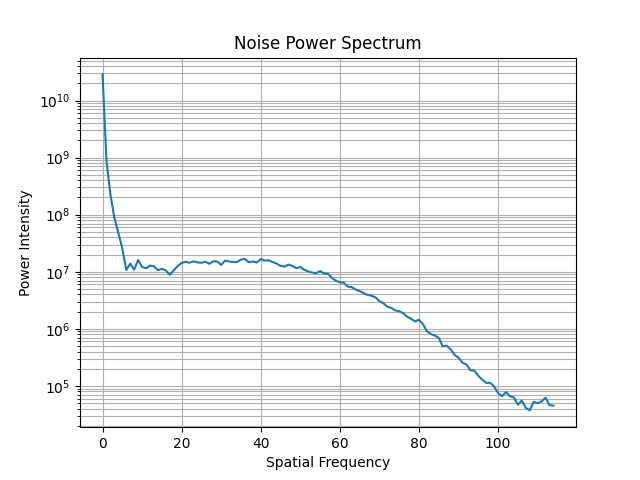
Advanced Use#
Using results_data#
Changed in version 3.0.
Using the catphan module in your own scripts? While the analysis results can be printed out,
if you intend on using them elsewhere (e.g. in an API), they can be accessed the easiest by using the results_data() method
which returns a CatPhanResult instance.
Note
While the pylinac tooling may change under the hood, this object should remain largely the same and/or expand. Thus, using this is more stable than accessing attrs directly.
Continuing from above:
data = mycbct.results_data()
data.catphan_model
data.ctp404.measured_slice_thickness_mm
# and more
# return as a dict
data_dict = mycbct.results_data(as_dict=True)
data_dict["ctp404"]["measured_slice_thickness_mm"]
...
Partial scans#
While the default behavior of pylinac is to analyze all modules in the scan (in fact it will error out if they aren’t),
the behavior can be customized. Pylinac always has to be aware of the CTP404 module as that’s the reference slice
for everything else. Thus, if the 404 is not in the scan you’re SOL. However, if one of the other modules is not present
you can remove or adjust its offset by subclassing and overloading the modules attr:
from pylinac import CatPhan504 # works for any of the other phantoms too
from pylinac.ct import CTP515, CTP486
class PartialCatPhan504(CatPhan504):
modules = {
CTP486: {"offset": -65},
CTP515: {"offset": -30},
# the CTP528 was omitted
}
ct = PartialCatPhan504.from_zip(...) # use like normal
Examining rMTF#
The rMTF can be calculated ad hoc like so. Note that CTP528 must be present (see above):
ct = ... # load a dataset like normal
ct.analyze()
ct.ctp528.mtf.relative_resolution(x=40) # get the rMTF (lp/mm) at 40% resolution
Customizing module locations#
Similar to partial scans, to modify the module location(s), overload the modules attr and edit the offset value.
The value is in mm:
from pylinac import CatPhan504 # works for any of the other phantoms too
from pylinac.ct import CTP515, CTP486, CTP528
# create custom catphan with module locations
class OffsetCatPhan504(CatPhan504):
modules = {
CTP486: {"offset": -60}, # normally -65
CTP528: {"offset": 30},
CTP515: {"offset": -25}, # normally -30
}
ct = OffsetCatPhan504.from_zip(...) # use like normal
Customizing Modules#
You can also customize modules themselves in v2.4+. Customization should always be done by subclassing an existing module and overloading the attributes. Then, pass in the new custom module into the parent CatPhan class. The easiest way to get started is copy the relevant attributes from the existing code.
As an example, let’s override the angles of the ROIs for CTP404.
from pylinac.ct import CatPhan504, CTP404CP504
# first, customize the module
class CustomCTP404(CTP404CP504):
roi_dist_mm = 58.7 # this is the default value; we repeat here because it's easy to copy from source
roi_radius_mm = 5 # ditto
roi_settings = {
"Air": {
"value": -1000,
"angle": -93, # changed 'angle' from -90 to -93
"distance": roi_dist_mm,
"radius": roi_radius_mm,
},
"PMP": {
"value": -196,
"angle": -122, # changed 'angle' from -120 to -122
"distance": roi_dist_mm,
"radius": roi_radius_mm,
},
# add other ROIs as appropriate
}
# then, pass to the CatPhan model
class CustomCP504(CatPhan504):
modules = {
CustomCTP404: {"offset": 0}
# add other modules here as appropriate
}
# use like normal
ct = CustomCP504(...)
Warning
If you overload the roi_settings or modules attributes, you are responsible for filling it out completely.
I.e. when you overload it’s not partial. In the above example if you want other CTP modules you must populate them.
Algorithm#
The CatPhan module is based on the tests and values given in the respective CatPhan manual. The algorithm works like such:
Allowances#
The images can be any size.
The phantom can have significant translation in all 3 directions.
The phantom can have significant roll and moderate yaw and pitch.
Restrictions#
Warning
Analysis can fail or give unreliable results if any Restriction is violated.
All of the modules defined in the
modulesattribute must be within the scan extent.Scan slices are not expected to overlap.
Warning
Overlapping slices are not generally a problem other than the slice thickness measurement. See the Slice Thickness section for how to override this to get a valid slice thickness in such a situation.
Pre-Analysis#
Determine image properties – Upon load, the image set is analyzed for its DICOM properties to determine mm/pixel spacing, rescale intercept and slope, manufacturer, etc.
Convert to HU – The entire image set is converted from its raw values to HU by applying the rescale intercept and slope which is contained in the DICOM properties.
Find the phantom z-location – Upon loading, all the images are scanned to determine where the HU linearity module (CTP404) is located. This is accomplished by examining each image slice and looking for 2 things:
If the CatPhan is in the image. At the edges of the scan this may not be true.
If a circular profile has characteristics like the CTP404 module. If the CatPhan is in the image, a circular profile is taken at the location where the HU linearity regions of interest are located. If the profile contains low, high, and lots of medium values then it is very likely the HU linearity module. All such slices are found and the median slice is set as the HU linearity module location. All other modules are located relative to this position.
Analysis#
Determine phantom roll – Precise knowledge of the ROIs to analyze is important, and small changes in rotation could invalidate automatic results. The roll of the phantom is determined by examining the HU module and converting to a binary image. The air holes are then located and the angle of the two holes determines the phantom roll.
Note
For each step below, the “module” analyzed is actually the mean, median, or maximum of 3 slices (+/-1 slice around and including the nominal slice) to ensure robust measurements. Also, for each step/phantom module, the phantom center is determined, which corrects for the phantom pitch and yaw.
Additionally, values tend to be lazy (computed only when asked for), thus the calculations listed may sometimes be performed only when asked for.
Determine HU linearity – The HU module (CTP404) contains several materials with different HU values. Using hardcoded angles (corrected for roll) and radius from the center of the phantom, circular ROIs are sampled which correspond to the HU material regions. The median pixel value of the ROI is the stated HU value. Nominal HU values are taken as the mean of the range given in the manual(s):
Note
As of v3.16, these can be overriden: Custom HU values.
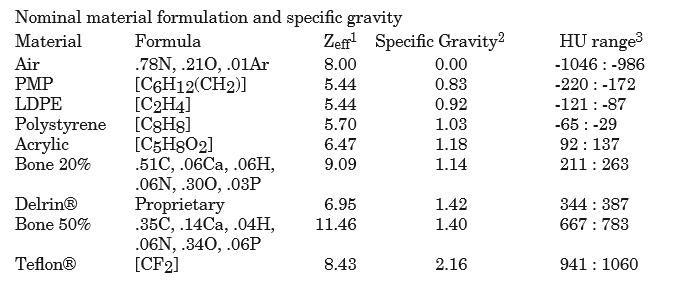
Determine HU uniformity – HU uniformity (CTP486) is calculated in a similar manner to HU linearity, but within the CTP486 module/slice.
Calculate Geometry/Scaling – The HU module (CTP404), besides HU materials, also contains several “nodes” which have an accurate spacing (50 mm apart). Again, using hardcoded but corrected angles, the area around the 4 nodes are sampled and then a threshold is applied which identifies the node within the ROI sample. The center of mass of the node is determined and then the space between nodes is calculated.
Calculate Spatial Resolution/MTF – The Spatial Resolution module (CTP528) contains 21 pairs of aluminum bars having varying thickness, which also corresponds to the thickness between the bars. One unique advantage of these bars is that they are all focused on and equally distant to the phantom center. This is taken advantage of by extracting a
CollapsedCircleProfileabout the line pairs. The peaks and valleys of the profile are located; peaks and valleys of each line pair are used to calculated the MTF. The relative MTF (i.e. normalized to the first line pair) is then calculated from these values. See Modulation Transfer Function.Calculate Low Contrast Resolution – Low contrast is inherently difficult to determine since detectability of humans is not simply contrast based. Pylinac’s analysis uses both the contrast value of the ROI as well as the ROI size to compute a “detectability” score. ROIs above the score are said to be “seen”, while those below are not seen. Only the 1.0% supra-slice ROIs are examined. Two background ROIs are sampled on either side of the ROI contrast set. See Visibility for equation details.
Calculate Slice Thickness – Slice thickness is measured by determining the FWHM of the wire ramps in the CTP404 module. A profile of the area around each wire ramp is taken, and the FWHM is determined from the profile. The profiles are averaged and the value is converted from pixels to mm and multiplied by 0.42 (Catphan manual “Scan Slice Geometry” section). Also see Slice Thickness.
Post-Analysis#
Test if values are within tolerance – For each module, the determined values are compared with the nominal values. If the difference between the two is below the specified tolerance then the module passes.
Troubleshooting#
First, check the general Troubleshooting section. Most problems in this module revolve around getting the data loaded.
If you’re having trouble getting your dataset in, make sure you’re loading the whole dataset. Also make sure you’ve scanned the whole phantom.
Make sure there are no external markers on the CatPhan (e.g. BBs), otherwise the localization algorithm will not be able to properly locate the phantom within the image.
Ensure that the FOV is large enough to encompass the entire phantom. If the scan is cutting off the phantom in any way it will not identify it.
The phantom should never touch the edge of an image, see above point.
Make sure you’re loading the right CatPhan class. I.e. using a CatPhan600 class on a CatPhan504 scan may result in errors or erroneous results.
API Documentation#
Main classes#
These are the classes a typical user may interface with.
- class pylinac.ct.CatPhan504(folderpath: str | Sequence[str] | Path | Sequence[Path] | Sequence[BytesIO], check_uid: bool = True, memory_efficient_mode: bool = False)[source]#
Bases:
CatPhanBaseA class for loading and analyzing CT DICOM files of a CatPhan 504. Can be from a CBCT or CT scanner Analyzes: Uniformity (CTP486), High-Contrast Spatial Resolution (CTP528), Image Scaling & HU Linearity (CTP404), and Low contrast (CTP515).
Parameters#
- folderpathstr, list of strings, or Path to folder
String that points to the CBCT image folder location.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- memory_efficient_modebool
Whether to use a memory efficient mode. If True, the DICOM stack will be loaded on demand rather than all at once. This will reduce the memory footprint but will be slower by ~25%. Default is False.
Raises#
- NotADirectoryError
If folder str passed is not a valid directory.
- FileNotFoundError
If no CT images are found in the folder
- static run_demo(show: bool = True)[source]#
Run the CBCT demo using high-quality head protocol images.
- analyze(hu_tolerance: int | float = 40, scaling_tolerance: int | float = 1, thickness_tolerance: int | float = 0.2, low_contrast_tolerance: int | float = 1, cnr_threshold: int | float = 15, zip_after: bool = False, contrast_method: str = 'Michelson', visibility_threshold: float = 0.15, thickness_slice_straddle: str | int = 'auto', expected_hu_values: dict[str, int | float] | None = None)#
Single-method full analysis of CBCT DICOM files.
Parameters#
- hu_toleranceint
The HU tolerance value for both HU uniformity and linearity.
- scaling_tolerancefloat, int
The scaling tolerance in mm of the geometric nodes on the HU linearity slice (CTP404 module).
- thickness_tolerancefloat, int
The tolerance of the thickness calculation in mm, based on the wire ramps in the CTP404 module.
Warning
Thickness accuracy degrades with image noise; i.e. low mAs images are less accurate.
- low_contrast_toleranceint
The number of low-contrast bubbles needed to be “seen” to pass.
- cnr_thresholdfloat, int
The threshold for “detecting” low-contrast image. See RTD for calculation info.
Deprecated since version 3.0: Use visibility parameter instead.
- zip_afterbool
If the CT images were not compressed before analysis and this is set to true, pylinac will compress the analyzed images into a ZIP archive.
- contrast_method
The contrast equation to use. See Low contrast.
- visibility_threshold
The threshold for detecting low-contrast ROIs. Use instead of
cnr_threshold. Follows the Rose equation. See Visibility.- thickness_slice_straddle
The number of extra slices on each side of the HU module slice to use for slice thickness determination. The rationale is that for thin slices the ramp FWHM can be very noisy. I.e. a 1mm slice might have a 100% variation with a low-mAs protocol. To account for this, slice thicknesses < 3.5mm have 1 slice added on either side of the HU module (so 3 total slices) and then averaged. The default is ‘auto’, which follows the above logic. Set to an integer to explicitly use a certain amount of padding. Typical values are 0, 1, and 2.
Warning
This is the padding on either side. So a value of 1 => 3 slices, 2 => 5 slices, 3 => 7 slices, etc.
- expected_hu_values
An optional dictionary of the expected HU values for the HU linearity module. The keys are the ROI names and the values are the expected HU values. If a key is not present or the parameter is None, the default values will be used.
- property catphan_size: float#
The expected size of the phantom in pixels, based on a 20cm wide phantom.
- find_origin_slice() int#
Using a brute force search of the images, find the median HU linearity slice.
This method walks through all the images and takes a collapsed circle profile where the HU linearity ROIs are. If the profile contains both low (<800) and high (>800) HU values and most values are the same (i.e. it’s not an artifact), then it can be assumed it is an HU linearity slice. The median of all applicable slices is the center of the HU slice.
Returns#
- int
The middle slice of the HU linearity module.
- find_phantom_axis()#
We fit all the center locations of the phantom across all slices to a 1D poly function instead of finding them individually for robustness.
Normally, each slice would be evaluated individually, but the RadMachine jig gets in the way of detecting the HU module (🤦♂️). To work around that in a backwards-compatible way we instead look at all the slices and if the phantom was detected, capture the phantom center. ALL the centers are then fitted to a 1D poly function and passed to the individual slices. This way, even if one slice is messed up (such as because of the phantom jig), the poly function is robust to give the real center based on all the other properly-located positions on the other slices.
- find_phantom_roll(func: Callable | None = None) float#
Determine the “roll” of the phantom.
This algorithm uses the two air bubbles in the HU slice and the resulting angle between them.
Parameters#
- func
A callable to sort the air ROIs.
Returns#
float : the angle of the phantom in degrees.
- classmethod from_demo_images()#
Construct a CBCT object from the demo images.
- classmethod from_url(url: str, check_uid: bool = True)#
Instantiate a CBCT object from a URL pointing to a .zip object.
Parameters#
- urlstr
URL pointing to a zip archive of CBCT images.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- classmethod from_zip(zip_file: str | ZipFile | BinaryIO, check_uid: bool = True, memory_efficient_mode: bool = False)#
Construct a CBCT object and pass the zip file.
Parameters#
- zip_filestr, ZipFile
Path to the zip file or a ZipFile object.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- memory_efficient_modebool
Whether to use a memory efficient mode. If True, the DICOM stack will be loaded on demand rather than all at once. This will reduce the memory footprint but will be slower by ~25%. Default is False.
Raises#
FileExistsError : If zip_file passed was not a legitimate zip file. FileNotFoundError : If no CT images are found in the folder
- localize() None#
Find the slice number of the catphan’s HU linearity module and roll angle
- property mm_per_pixel: float#
The millimeters per pixel of the DICOM images.
- property num_images: int#
The number of images loaded.
- plot_analyzed_image(show: bool = True, **plt_kwargs) None#
Plot the images used in the calculation and summary data.
Parameters#
- showbool
Whether to plot the image or not.
- plt_kwargsdict
Keyword args passed to the plt.figure() method. Allows one to set things like figure size.
- plot_analyzed_subimage(subimage: str = 'hu', delta: bool = True, show: bool = True) Figure | None#
Plot a specific component of the CBCT analysis.
Parameters#
- subimage{‘hu’, ‘un’, ‘sp’, ‘lc’, ‘mtf’, ‘lin’, ‘prof’, ‘side’}
The subcomponent to plot. Values must contain one of the following letter combinations. E.g.
linearity,linear, andlinwill all draw the HU linearity values.hudraws the HU linearity image.undraws the HU uniformity image.spdraws the Spatial Resolution image.lcdraws the Low Contrast image (if applicable).mtfdraws the RMTF plot.lindraws the HU linearity values. Used withdelta.profdraws the HU uniformity profiles.sidedraws the side view of the phantom with lines of the module locations.
- deltabool
Only for use with
lin. Whether to plot the HU delta or actual values.- showbool
Whether to actually show the plot.
- plot_side_view(axis: Axes) None#
Plot a view of the scan from the side with lines showing detected module positions
- publish_pdf(filename: str | Path, notes: str | None = None, open_file: bool = False, metadata: dict | None = None, logo: Path | str | None = None) None#
Publish (print) a PDF containing the analysis and quantitative results.
Parameters#
- filename(str, file-like object}
The file to write the results to.
- notesstr, list of strings
Text; if str, prints single line. If list of strings, each list item is printed on its own line.
- open_filebool
Whether to open the file using the default program after creation.
- metadatadict
Extra data to be passed and shown in the PDF. The key and value will be shown with a colon. E.g. passing {‘Author’: ‘James’, ‘Unit’: ‘TrueBeam’} would result in text in the PDF like: ————– Author: James Unit: TrueBeam ————–
- logo: Path, str
A custom logo to use in the PDF report. If nothing is passed, the default pylinac logo is used.
- refine_origin_slice(initial_slice_num: int) int#
Apply a refinement to the origin slice. This was added to handle the catphan 604 at least due to variations in the length of the HU plugs.
- results(as_list: bool = False) str | list[list[str]]#
Return the results of the analysis as a string. Use with print().
Parameters#
- as_listbool
Whether to return as a list of list of strings vs single string. Pretty much for internal usage.
- results_data(as_dict: bool = False, as_json: bool = False) T | dict | str#
Present the results data and metadata as a dataclass, dict, or tuple. The default return type is a dataclass.
Parameters#
- as_dictbool
If True, return the results as a dictionary.
- as_jsonbool
If True, return the results as a JSON string. Cannot be True if as_dict is True.
- save_analyzed_image(filename: str | Path | BinaryIO, **kwargs) None#
Save the analyzed summary plot.
Parameters#
- filenamestr, file object
The name of the file to save the image to.
- kwargs :
Any valid matplotlib kwargs.
- save_analyzed_subimage(filename: str | BinaryIO, subimage: str = 'hu', delta: bool = True, **kwargs) Figure | None#
Save a component image to file.
Parameters#
- filenamestr, file object
The file to write the image to.
- subimagestr
See
plot_analyzed_subimage()for parameter info.- deltabool
Only for use with
lin. Whether to plot the HU delta or actual values.
- class pylinac.ct.CatPhan503(folderpath: str | Sequence[str] | Path | Sequence[Path] | Sequence[BytesIO], check_uid: bool = True, memory_efficient_mode: bool = False)[source]#
Bases:
CatPhanBaseA class for loading and analyzing CT DICOM files of a CatPhan 503. Analyzes: Uniformity (CTP486), High-Contrast Spatial Resolution (CTP528), Image Scaling & HU Linearity (CTP404).
Parameters#
- folderpathstr, list of strings, or Path to folder
String that points to the CBCT image folder location.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- memory_efficient_modebool
Whether to use a memory efficient mode. If True, the DICOM stack will be loaded on demand rather than all at once. This will reduce the memory footprint but will be slower by ~25%. Default is False.
Raises#
- NotADirectoryError
If folder str passed is not a valid directory.
- FileNotFoundError
If no CT images are found in the folder
- static run_demo(show: bool = True)[source]#
Run the CBCT demo using high-quality head protocol images.
- analyze(hu_tolerance: int | float = 40, scaling_tolerance: int | float = 1, thickness_tolerance: int | float = 0.2, low_contrast_tolerance: int | float = 1, cnr_threshold: int | float = 15, zip_after: bool = False, contrast_method: str = 'Michelson', visibility_threshold: float = 0.15, thickness_slice_straddle: str | int = 'auto', expected_hu_values: dict[str, int | float] | None = None)#
Single-method full analysis of CBCT DICOM files.
Parameters#
- hu_toleranceint
The HU tolerance value for both HU uniformity and linearity.
- scaling_tolerancefloat, int
The scaling tolerance in mm of the geometric nodes on the HU linearity slice (CTP404 module).
- thickness_tolerancefloat, int
The tolerance of the thickness calculation in mm, based on the wire ramps in the CTP404 module.
Warning
Thickness accuracy degrades with image noise; i.e. low mAs images are less accurate.
- low_contrast_toleranceint
The number of low-contrast bubbles needed to be “seen” to pass.
- cnr_thresholdfloat, int
The threshold for “detecting” low-contrast image. See RTD for calculation info.
Deprecated since version 3.0: Use visibility parameter instead.
- zip_afterbool
If the CT images were not compressed before analysis and this is set to true, pylinac will compress the analyzed images into a ZIP archive.
- contrast_method
The contrast equation to use. See Low contrast.
- visibility_threshold
The threshold for detecting low-contrast ROIs. Use instead of
cnr_threshold. Follows the Rose equation. See Visibility.- thickness_slice_straddle
The number of extra slices on each side of the HU module slice to use for slice thickness determination. The rationale is that for thin slices the ramp FWHM can be very noisy. I.e. a 1mm slice might have a 100% variation with a low-mAs protocol. To account for this, slice thicknesses < 3.5mm have 1 slice added on either side of the HU module (so 3 total slices) and then averaged. The default is ‘auto’, which follows the above logic. Set to an integer to explicitly use a certain amount of padding. Typical values are 0, 1, and 2.
Warning
This is the padding on either side. So a value of 1 => 3 slices, 2 => 5 slices, 3 => 7 slices, etc.
- expected_hu_values
An optional dictionary of the expected HU values for the HU linearity module. The keys are the ROI names and the values are the expected HU values. If a key is not present or the parameter is None, the default values will be used.
- property catphan_size: float#
The expected size of the phantom in pixels, based on a 20cm wide phantom.
- find_origin_slice() int#
Using a brute force search of the images, find the median HU linearity slice.
This method walks through all the images and takes a collapsed circle profile where the HU linearity ROIs are. If the profile contains both low (<800) and high (>800) HU values and most values are the same (i.e. it’s not an artifact), then it can be assumed it is an HU linearity slice. The median of all applicable slices is the center of the HU slice.
Returns#
- int
The middle slice of the HU linearity module.
- find_phantom_axis()#
We fit all the center locations of the phantom across all slices to a 1D poly function instead of finding them individually for robustness.
Normally, each slice would be evaluated individually, but the RadMachine jig gets in the way of detecting the HU module (🤦♂️). To work around that in a backwards-compatible way we instead look at all the slices and if the phantom was detected, capture the phantom center. ALL the centers are then fitted to a 1D poly function and passed to the individual slices. This way, even if one slice is messed up (such as because of the phantom jig), the poly function is robust to give the real center based on all the other properly-located positions on the other slices.
- find_phantom_roll(func: Callable | None = None) float#
Determine the “roll” of the phantom.
This algorithm uses the two air bubbles in the HU slice and the resulting angle between them.
Parameters#
- func
A callable to sort the air ROIs.
Returns#
float : the angle of the phantom in degrees.
- classmethod from_demo_images()#
Construct a CBCT object from the demo images.
- classmethod from_url(url: str, check_uid: bool = True)#
Instantiate a CBCT object from a URL pointing to a .zip object.
Parameters#
- urlstr
URL pointing to a zip archive of CBCT images.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- classmethod from_zip(zip_file: str | ZipFile | BinaryIO, check_uid: bool = True, memory_efficient_mode: bool = False)#
Construct a CBCT object and pass the zip file.
Parameters#
- zip_filestr, ZipFile
Path to the zip file or a ZipFile object.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- memory_efficient_modebool
Whether to use a memory efficient mode. If True, the DICOM stack will be loaded on demand rather than all at once. This will reduce the memory footprint but will be slower by ~25%. Default is False.
Raises#
FileExistsError : If zip_file passed was not a legitimate zip file. FileNotFoundError : If no CT images are found in the folder
- localize() None#
Find the slice number of the catphan’s HU linearity module and roll angle
- property mm_per_pixel: float#
The millimeters per pixel of the DICOM images.
- property num_images: int#
The number of images loaded.
- plot_analyzed_image(show: bool = True, **plt_kwargs) None#
Plot the images used in the calculation and summary data.
Parameters#
- showbool
Whether to plot the image or not.
- plt_kwargsdict
Keyword args passed to the plt.figure() method. Allows one to set things like figure size.
- plot_analyzed_subimage(subimage: str = 'hu', delta: bool = True, show: bool = True) Figure | None#
Plot a specific component of the CBCT analysis.
Parameters#
- subimage{‘hu’, ‘un’, ‘sp’, ‘lc’, ‘mtf’, ‘lin’, ‘prof’, ‘side’}
The subcomponent to plot. Values must contain one of the following letter combinations. E.g.
linearity,linear, andlinwill all draw the HU linearity values.hudraws the HU linearity image.undraws the HU uniformity image.spdraws the Spatial Resolution image.lcdraws the Low Contrast image (if applicable).mtfdraws the RMTF plot.lindraws the HU linearity values. Used withdelta.profdraws the HU uniformity profiles.sidedraws the side view of the phantom with lines of the module locations.
- deltabool
Only for use with
lin. Whether to plot the HU delta or actual values.- showbool
Whether to actually show the plot.
- plot_side_view(axis: Axes) None#
Plot a view of the scan from the side with lines showing detected module positions
- publish_pdf(filename: str | Path, notes: str | None = None, open_file: bool = False, metadata: dict | None = None, logo: Path | str | None = None) None#
Publish (print) a PDF containing the analysis and quantitative results.
Parameters#
- filename(str, file-like object}
The file to write the results to.
- notesstr, list of strings
Text; if str, prints single line. If list of strings, each list item is printed on its own line.
- open_filebool
Whether to open the file using the default program after creation.
- metadatadict
Extra data to be passed and shown in the PDF. The key and value will be shown with a colon. E.g. passing {‘Author’: ‘James’, ‘Unit’: ‘TrueBeam’} would result in text in the PDF like: ————– Author: James Unit: TrueBeam ————–
- logo: Path, str
A custom logo to use in the PDF report. If nothing is passed, the default pylinac logo is used.
- refine_origin_slice(initial_slice_num: int) int#
Apply a refinement to the origin slice. This was added to handle the catphan 604 at least due to variations in the length of the HU plugs.
- results(as_list: bool = False) str | list[list[str]]#
Return the results of the analysis as a string. Use with print().
Parameters#
- as_listbool
Whether to return as a list of list of strings vs single string. Pretty much for internal usage.
- results_data(as_dict: bool = False, as_json: bool = False) T | dict | str#
Present the results data and metadata as a dataclass, dict, or tuple. The default return type is a dataclass.
Parameters#
- as_dictbool
If True, return the results as a dictionary.
- as_jsonbool
If True, return the results as a JSON string. Cannot be True if as_dict is True.
- save_analyzed_image(filename: str | Path | BinaryIO, **kwargs) None#
Save the analyzed summary plot.
Parameters#
- filenamestr, file object
The name of the file to save the image to.
- kwargs :
Any valid matplotlib kwargs.
- save_analyzed_subimage(filename: str | BinaryIO, subimage: str = 'hu', delta: bool = True, **kwargs) Figure | None#
Save a component image to file.
Parameters#
- filenamestr, file object
The file to write the image to.
- subimagestr
See
plot_analyzed_subimage()for parameter info.- deltabool
Only for use with
lin. Whether to plot the HU delta or actual values.
- class pylinac.ct.CatPhan600(folderpath: str | Sequence[str] | Path | Sequence[Path] | Sequence[BytesIO], check_uid: bool = True, memory_efficient_mode: bool = False)[source]#
Bases:
CatPhanBaseA class for loading and analyzing CT DICOM files of a CatPhan 600. Analyzes: Uniformity (CTP486), High-Contrast Spatial Resolution (CTP528), Image Scaling & HU Linearity (CTP404), and Low contrast (CTP515).
Parameters#
- folderpathstr, list of strings, or Path to folder
String that points to the CBCT image folder location.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- memory_efficient_modebool
Whether to use a memory efficient mode. If True, the DICOM stack will be loaded on demand rather than all at once. This will reduce the memory footprint but will be slower by ~25%. Default is False.
Raises#
- NotADirectoryError
If folder str passed is not a valid directory.
- FileNotFoundError
If no CT images are found in the folder
- find_phantom_roll(func: Callable | None = None) float[source]#
With the CatPhan 600, we have to consider that the top air ROI has a water vial in it (see pg 12 of the manual). If so, the top air ROI won’t be detected. Rather, the default algorithm will find the bottom air ROI and teflon to the left. It may also find the top air ROI if the water vial isn’t there. We use the below lambda to select the bottom air and teflon ROIs consistently. These two ROIs are at 75 degrees from cardinal. We thus offset the default outcome by 75.
- analyze(hu_tolerance: int | float = 40, scaling_tolerance: int | float = 1, thickness_tolerance: int | float = 0.2, low_contrast_tolerance: int | float = 1, cnr_threshold: int | float = 15, zip_after: bool = False, contrast_method: str = 'Michelson', visibility_threshold: float = 0.15, thickness_slice_straddle: str | int = 'auto', expected_hu_values: dict[str, int | float] | None = None)#
Single-method full analysis of CBCT DICOM files.
Parameters#
- hu_toleranceint
The HU tolerance value for both HU uniformity and linearity.
- scaling_tolerancefloat, int
The scaling tolerance in mm of the geometric nodes on the HU linearity slice (CTP404 module).
- thickness_tolerancefloat, int
The tolerance of the thickness calculation in mm, based on the wire ramps in the CTP404 module.
Warning
Thickness accuracy degrades with image noise; i.e. low mAs images are less accurate.
- low_contrast_toleranceint
The number of low-contrast bubbles needed to be “seen” to pass.
- cnr_thresholdfloat, int
The threshold for “detecting” low-contrast image. See RTD for calculation info.
Deprecated since version 3.0: Use visibility parameter instead.
- zip_afterbool
If the CT images were not compressed before analysis and this is set to true, pylinac will compress the analyzed images into a ZIP archive.
- contrast_method
The contrast equation to use. See Low contrast.
- visibility_threshold
The threshold for detecting low-contrast ROIs. Use instead of
cnr_threshold. Follows the Rose equation. See Visibility.- thickness_slice_straddle
The number of extra slices on each side of the HU module slice to use for slice thickness determination. The rationale is that for thin slices the ramp FWHM can be very noisy. I.e. a 1mm slice might have a 100% variation with a low-mAs protocol. To account for this, slice thicknesses < 3.5mm have 1 slice added on either side of the HU module (so 3 total slices) and then averaged. The default is ‘auto’, which follows the above logic. Set to an integer to explicitly use a certain amount of padding. Typical values are 0, 1, and 2.
Warning
This is the padding on either side. So a value of 1 => 3 slices, 2 => 5 slices, 3 => 7 slices, etc.
- expected_hu_values
An optional dictionary of the expected HU values for the HU linearity module. The keys are the ROI names and the values are the expected HU values. If a key is not present or the parameter is None, the default values will be used.
- property catphan_size: float#
The expected size of the phantom in pixels, based on a 20cm wide phantom.
- find_origin_slice() int#
Using a brute force search of the images, find the median HU linearity slice.
This method walks through all the images and takes a collapsed circle profile where the HU linearity ROIs are. If the profile contains both low (<800) and high (>800) HU values and most values are the same (i.e. it’s not an artifact), then it can be assumed it is an HU linearity slice. The median of all applicable slices is the center of the HU slice.
Returns#
- int
The middle slice of the HU linearity module.
- find_phantom_axis()#
We fit all the center locations of the phantom across all slices to a 1D poly function instead of finding them individually for robustness.
Normally, each slice would be evaluated individually, but the RadMachine jig gets in the way of detecting the HU module (🤦♂️). To work around that in a backwards-compatible way we instead look at all the slices and if the phantom was detected, capture the phantom center. ALL the centers are then fitted to a 1D poly function and passed to the individual slices. This way, even if one slice is messed up (such as because of the phantom jig), the poly function is robust to give the real center based on all the other properly-located positions on the other slices.
- classmethod from_demo_images()#
Construct a CBCT object from the demo images.
- classmethod from_url(url: str, check_uid: bool = True)#
Instantiate a CBCT object from a URL pointing to a .zip object.
Parameters#
- urlstr
URL pointing to a zip archive of CBCT images.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- classmethod from_zip(zip_file: str | ZipFile | BinaryIO, check_uid: bool = True, memory_efficient_mode: bool = False)#
Construct a CBCT object and pass the zip file.
Parameters#
- zip_filestr, ZipFile
Path to the zip file or a ZipFile object.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- memory_efficient_modebool
Whether to use a memory efficient mode. If True, the DICOM stack will be loaded on demand rather than all at once. This will reduce the memory footprint but will be slower by ~25%. Default is False.
Raises#
FileExistsError : If zip_file passed was not a legitimate zip file. FileNotFoundError : If no CT images are found in the folder
- localize() None#
Find the slice number of the catphan’s HU linearity module and roll angle
- property mm_per_pixel: float#
The millimeters per pixel of the DICOM images.
- property num_images: int#
The number of images loaded.
- plot_analyzed_image(show: bool = True, **plt_kwargs) None#
Plot the images used in the calculation and summary data.
Parameters#
- showbool
Whether to plot the image or not.
- plt_kwargsdict
Keyword args passed to the plt.figure() method. Allows one to set things like figure size.
- plot_analyzed_subimage(subimage: str = 'hu', delta: bool = True, show: bool = True) Figure | None#
Plot a specific component of the CBCT analysis.
Parameters#
- subimage{‘hu’, ‘un’, ‘sp’, ‘lc’, ‘mtf’, ‘lin’, ‘prof’, ‘side’}
The subcomponent to plot. Values must contain one of the following letter combinations. E.g.
linearity,linear, andlinwill all draw the HU linearity values.hudraws the HU linearity image.undraws the HU uniformity image.spdraws the Spatial Resolution image.lcdraws the Low Contrast image (if applicable).mtfdraws the RMTF plot.lindraws the HU linearity values. Used withdelta.profdraws the HU uniformity profiles.sidedraws the side view of the phantom with lines of the module locations.
- deltabool
Only for use with
lin. Whether to plot the HU delta or actual values.- showbool
Whether to actually show the plot.
- plot_side_view(axis: Axes) None#
Plot a view of the scan from the side with lines showing detected module positions
- publish_pdf(filename: str | Path, notes: str | None = None, open_file: bool = False, metadata: dict | None = None, logo: Path | str | None = None) None#
Publish (print) a PDF containing the analysis and quantitative results.
Parameters#
- filename(str, file-like object}
The file to write the results to.
- notesstr, list of strings
Text; if str, prints single line. If list of strings, each list item is printed on its own line.
- open_filebool
Whether to open the file using the default program after creation.
- metadatadict
Extra data to be passed and shown in the PDF. The key and value will be shown with a colon. E.g. passing {‘Author’: ‘James’, ‘Unit’: ‘TrueBeam’} would result in text in the PDF like: ————– Author: James Unit: TrueBeam ————–
- logo: Path, str
A custom logo to use in the PDF report. If nothing is passed, the default pylinac logo is used.
- refine_origin_slice(initial_slice_num: int) int#
Apply a refinement to the origin slice. This was added to handle the catphan 604 at least due to variations in the length of the HU plugs.
- results(as_list: bool = False) str | list[list[str]]#
Return the results of the analysis as a string. Use with print().
Parameters#
- as_listbool
Whether to return as a list of list of strings vs single string. Pretty much for internal usage.
- results_data(as_dict: bool = False, as_json: bool = False) T | dict | str#
Present the results data and metadata as a dataclass, dict, or tuple. The default return type is a dataclass.
Parameters#
- as_dictbool
If True, return the results as a dictionary.
- as_jsonbool
If True, return the results as a JSON string. Cannot be True if as_dict is True.
- save_analyzed_image(filename: str | Path | BinaryIO, **kwargs) None#
Save the analyzed summary plot.
Parameters#
- filenamestr, file object
The name of the file to save the image to.
- kwargs :
Any valid matplotlib kwargs.
- save_analyzed_subimage(filename: str | BinaryIO, subimage: str = 'hu', delta: bool = True, **kwargs) Figure | None#
Save a component image to file.
Parameters#
- filenamestr, file object
The file to write the image to.
- subimagestr
See
plot_analyzed_subimage()for parameter info.- deltabool
Only for use with
lin. Whether to plot the HU delta or actual values.
- class pylinac.ct.CatPhan604(folderpath: str | Sequence[str] | Path | Sequence[Path] | Sequence[BytesIO], check_uid: bool = True, memory_efficient_mode: bool = False)[source]#
Bases:
CatPhanBaseA class for loading and analyzing CT DICOM files of a CatPhan 604. Can be from a CBCT or CT scanner Analyzes: Uniformity (CTP486), High-Contrast Spatial Resolution (CTP528), Image Scaling & HU Linearity (CTP404), and Low contrast (CTP515).
Parameters#
- folderpathstr, list of strings, or Path to folder
String that points to the CBCT image folder location.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- memory_efficient_modebool
Whether to use a memory efficient mode. If True, the DICOM stack will be loaded on demand rather than all at once. This will reduce the memory footprint but will be slower by ~25%. Default is False.
Raises#
- NotADirectoryError
If folder str passed is not a valid directory.
- FileNotFoundError
If no CT images are found in the folder
- static run_demo(show: bool = True)[source]#
Run the CBCT demo using high-quality head protocol images.
- refine_origin_slice(initial_slice_num: int) int[source]#
The HU plugs are longer than the ‘wire section’. This applies a refinement to find the slice that has the least angle between the centers of the left and right wires.
Starting with the initial slice, go +/- 5 slices to find the slice with the least angle between the left and right wires.
This suffers from a weakness that the roll is not yet determined. This will thus return the slice that has the least ABSOLUTE roll. If the phantom has an inherent roll, this will not be detected and may be off by a slice or so. Given the angle of the wire, the error would be small and likely only 1-2 slices max.
- analyze(hu_tolerance: int | float = 40, scaling_tolerance: int | float = 1, thickness_tolerance: int | float = 0.2, low_contrast_tolerance: int | float = 1, cnr_threshold: int | float = 15, zip_after: bool = False, contrast_method: str = 'Michelson', visibility_threshold: float = 0.15, thickness_slice_straddle: str | int = 'auto', expected_hu_values: dict[str, int | float] | None = None)#
Single-method full analysis of CBCT DICOM files.
Parameters#
- hu_toleranceint
The HU tolerance value for both HU uniformity and linearity.
- scaling_tolerancefloat, int
The scaling tolerance in mm of the geometric nodes on the HU linearity slice (CTP404 module).
- thickness_tolerancefloat, int
The tolerance of the thickness calculation in mm, based on the wire ramps in the CTP404 module.
Warning
Thickness accuracy degrades with image noise; i.e. low mAs images are less accurate.
- low_contrast_toleranceint
The number of low-contrast bubbles needed to be “seen” to pass.
- cnr_thresholdfloat, int
The threshold for “detecting” low-contrast image. See RTD for calculation info.
Deprecated since version 3.0: Use visibility parameter instead.
- zip_afterbool
If the CT images were not compressed before analysis and this is set to true, pylinac will compress the analyzed images into a ZIP archive.
- contrast_method
The contrast equation to use. See Low contrast.
- visibility_threshold
The threshold for detecting low-contrast ROIs. Use instead of
cnr_threshold. Follows the Rose equation. See Visibility.- thickness_slice_straddle
The number of extra slices on each side of the HU module slice to use for slice thickness determination. The rationale is that for thin slices the ramp FWHM can be very noisy. I.e. a 1mm slice might have a 100% variation with a low-mAs protocol. To account for this, slice thicknesses < 3.5mm have 1 slice added on either side of the HU module (so 3 total slices) and then averaged. The default is ‘auto’, which follows the above logic. Set to an integer to explicitly use a certain amount of padding. Typical values are 0, 1, and 2.
Warning
This is the padding on either side. So a value of 1 => 3 slices, 2 => 5 slices, 3 => 7 slices, etc.
- expected_hu_values
An optional dictionary of the expected HU values for the HU linearity module. The keys are the ROI names and the values are the expected HU values. If a key is not present or the parameter is None, the default values will be used.
- property catphan_size: float#
The expected size of the phantom in pixels, based on a 20cm wide phantom.
- find_origin_slice() int#
Using a brute force search of the images, find the median HU linearity slice.
This method walks through all the images and takes a collapsed circle profile where the HU linearity ROIs are. If the profile contains both low (<800) and high (>800) HU values and most values are the same (i.e. it’s not an artifact), then it can be assumed it is an HU linearity slice. The median of all applicable slices is the center of the HU slice.
Returns#
- int
The middle slice of the HU linearity module.
- find_phantom_axis()#
We fit all the center locations of the phantom across all slices to a 1D poly function instead of finding them individually for robustness.
Normally, each slice would be evaluated individually, but the RadMachine jig gets in the way of detecting the HU module (🤦♂️). To work around that in a backwards-compatible way we instead look at all the slices and if the phantom was detected, capture the phantom center. ALL the centers are then fitted to a 1D poly function and passed to the individual slices. This way, even if one slice is messed up (such as because of the phantom jig), the poly function is robust to give the real center based on all the other properly-located positions on the other slices.
- find_phantom_roll(func: Callable | None = None) float#
Determine the “roll” of the phantom.
This algorithm uses the two air bubbles in the HU slice and the resulting angle between them.
Parameters#
- func
A callable to sort the air ROIs.
Returns#
float : the angle of the phantom in degrees.
- classmethod from_demo_images()#
Construct a CBCT object from the demo images.
- classmethod from_url(url: str, check_uid: bool = True)#
Instantiate a CBCT object from a URL pointing to a .zip object.
Parameters#
- urlstr
URL pointing to a zip archive of CBCT images.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- classmethod from_zip(zip_file: str | ZipFile | BinaryIO, check_uid: bool = True, memory_efficient_mode: bool = False)#
Construct a CBCT object and pass the zip file.
Parameters#
- zip_filestr, ZipFile
Path to the zip file or a ZipFile object.
- check_uidbool
Whether to enforce raising an error if more than one UID is found in the dataset.
- memory_efficient_modebool
Whether to use a memory efficient mode. If True, the DICOM stack will be loaded on demand rather than all at once. This will reduce the memory footprint but will be slower by ~25%. Default is False.
Raises#
FileExistsError : If zip_file passed was not a legitimate zip file. FileNotFoundError : If no CT images are found in the folder
- localize() None#
Find the slice number of the catphan’s HU linearity module and roll angle
- property mm_per_pixel: float#
The millimeters per pixel of the DICOM images.
- property num_images: int#
The number of images loaded.
- plot_analyzed_image(show: bool = True, **plt_kwargs) None#
Plot the images used in the calculation and summary data.
Parameters#
- showbool
Whether to plot the image or not.
- plt_kwargsdict
Keyword args passed to the plt.figure() method. Allows one to set things like figure size.
- plot_analyzed_subimage(subimage: str = 'hu', delta: bool = True, show: bool = True) Figure | None#
Plot a specific component of the CBCT analysis.
Parameters#
- subimage{‘hu’, ‘un’, ‘sp’, ‘lc’, ‘mtf’, ‘lin’, ‘prof’, ‘side’}
The subcomponent to plot. Values must contain one of the following letter combinations. E.g.
linearity,linear, andlinwill all draw the HU linearity values.hudraws the HU linearity image.undraws the HU uniformity image.spdraws the Spatial Resolution image.lcdraws the Low Contrast image (if applicable).mtfdraws the RMTF plot.lindraws the HU linearity values. Used withdelta.profdraws the HU uniformity profiles.sidedraws the side view of the phantom with lines of the module locations.
- deltabool
Only for use with
lin. Whether to plot the HU delta or actual values.- showbool
Whether to actually show the plot.
- plot_side_view(axis: Axes) None#
Plot a view of the scan from the side with lines showing detected module positions
- publish_pdf(filename: str | Path, notes: str | None = None, open_file: bool = False, metadata: dict | None = None, logo: Path | str | None = None) None#
Publish (print) a PDF containing the analysis and quantitative results.
Parameters#
- filename(str, file-like object}
The file to write the results to.
- notesstr, list of strings
Text; if str, prints single line. If list of strings, each list item is printed on its own line.
- open_filebool
Whether to open the file using the default program after creation.
- metadatadict
Extra data to be passed and shown in the PDF. The key and value will be shown with a colon. E.g. passing {‘Author’: ‘James’, ‘Unit’: ‘TrueBeam’} would result in text in the PDF like: ————– Author: James Unit: TrueBeam ————–
- logo: Path, str
A custom logo to use in the PDF report. If nothing is passed, the default pylinac logo is used.
- results(as_list: bool = False) str | list[list[str]]#
Return the results of the analysis as a string. Use with print().
Parameters#
- as_listbool
Whether to return as a list of list of strings vs single string. Pretty much for internal usage.
- results_data(as_dict: bool = False, as_json: bool = False) T | dict | str#
Present the results data and metadata as a dataclass, dict, or tuple. The default return type is a dataclass.
Parameters#
- as_dictbool
If True, return the results as a dictionary.
- as_jsonbool
If True, return the results as a JSON string. Cannot be True if as_dict is True.
- save_analyzed_image(filename: str | Path | BinaryIO, **kwargs) None#
Save the analyzed summary plot.
Parameters#
- filenamestr, file object
The name of the file to save the image to.
- kwargs :
Any valid matplotlib kwargs.
- save_analyzed_subimage(filename: str | BinaryIO, subimage: str = 'hu', delta: bool = True, **kwargs) Figure | None#
Save a component image to file.
Parameters#
- filenamestr, file object
The file to write the image to.
- subimagestr
See
plot_analyzed_subimage()for parameter info.- deltabool
Only for use with
lin. Whether to plot the HU delta or actual values.
- class pylinac.ct.CatphanResult(*, pylinac_version: str = '3.22.0', date_of_analysis: datetime = None, catphan_model: str, catphan_roll_deg: float, origin_slice: int, num_images: int, ctp404: CTP404Result, ctp486: CTP486Result | None = None, ctp528: CTP528Result | None = None, ctp515: CTP515Result | None = None)[source]#
Bases:
ResultBaseThis class should not be called directly. It is returned by the
results_data()method. It is a dataclass under the hood and thus comes with all the dunder magic.Use the following attributes as normal class attributes.
Create a new model by parsing and validating input data from keyword arguments.
Raises [ValidationError][pydantic_core.ValidationError] if the input data cannot be validated to form a valid model.
self is explicitly positional-only to allow self as a field name.
- catphan_model: str#
- catphan_roll_deg: float#
- origin_slice: int#
- num_images: int#
- ctp404: CTP404Result#
- ctp486: CTP486Result | None#
- ctp528: CTP528Result | None#
- ctp515: CTP515Result | None#
- model_computed_fields: ClassVar[dict[str, ComputedFieldInfo]] = {}#
A dictionary of computed field names and their corresponding ComputedFieldInfo objects.
- model_config: ClassVar[ConfigDict] = {'arbitrary_types_allowed': True}#
Configuration for the model, should be a dictionary conforming to [ConfigDict][pydantic.config.ConfigDict].
- model_fields: ClassVar[dict[str, FieldInfo]] = {'catphan_model': FieldInfo(annotation=str, required=True), 'catphan_roll_deg': FieldInfo(annotation=float, required=True), 'ctp404': FieldInfo(annotation=CTP404Result, required=True), 'ctp486': FieldInfo(annotation=Union[CTP486Result, NoneType], required=False, default=None), 'ctp515': FieldInfo(annotation=Union[CTP515Result, NoneType], required=False, default=None), 'ctp528': FieldInfo(annotation=Union[CTP528Result, NoneType], required=False, default=None), 'date_of_analysis': FieldInfo(annotation=datetime, required=False, default_factory=builtin_function_or_method), 'num_images': FieldInfo(annotation=int, required=True), 'origin_slice': FieldInfo(annotation=int, required=True), 'pylinac_version': FieldInfo(annotation=str, required=False, default='3.22.0')}#
Metadata about the fields defined on the model, mapping of field names to [FieldInfo][pydantic.fields.FieldInfo].
This replaces Model.__fields__ from Pydantic V1.
- class pylinac.ct.CTP404Result(*, offset: int, low_contrast_visibility: float, thickness_passed: bool, measured_slice_thickness_mm: float, thickness_num_slices_combined: int, geometry_passed: bool, avg_line_distance_mm: float, line_distances_mm: list[float], hu_linearity_passed: bool, hu_tolerance: float, hu_rois: dict)[source]#
Bases:
BaseModelThis class should not be called directly. It is returned by the
results_data()method. It is a dataclass under the hood and thus comes with all the dunder magic.Use the following attributes as normal class attributes.
Create a new model by parsing and validating input data from keyword arguments.
Raises [ValidationError][pydantic_core.ValidationError] if the input data cannot be validated to form a valid model.
self is explicitly positional-only to allow self as a field name.
- offset: int#
- low_contrast_visibility: float#
- thickness_passed: bool#
- measured_slice_thickness_mm: float#
- thickness_num_slices_combined: int#
- geometry_passed: bool#
- avg_line_distance_mm: float#
- line_distances_mm: list[float]#
- hu_linearity_passed: bool#
- hu_tolerance: float#
- hu_rois: dict#
- model_computed_fields: ClassVar[dict[str, ComputedFieldInfo]] = {}#
A dictionary of computed field names and their corresponding ComputedFieldInfo objects.
- model_config: ClassVar[ConfigDict] = {}#
Configuration for the model, should be a dictionary conforming to [ConfigDict][pydantic.config.ConfigDict].
- model_fields: ClassVar[dict[str, FieldInfo]] = {'avg_line_distance_mm': FieldInfo(annotation=float, required=True), 'geometry_passed': FieldInfo(annotation=bool, required=True), 'hu_linearity_passed': FieldInfo(annotation=bool, required=True), 'hu_rois': FieldInfo(annotation=dict, required=True), 'hu_tolerance': FieldInfo(annotation=float, required=True), 'line_distances_mm': FieldInfo(annotation=list[float], required=True), 'low_contrast_visibility': FieldInfo(annotation=float, required=True), 'measured_slice_thickness_mm': FieldInfo(annotation=float, required=True), 'offset': FieldInfo(annotation=int, required=True), 'thickness_num_slices_combined': FieldInfo(annotation=int, required=True), 'thickness_passed': FieldInfo(annotation=bool, required=True)}#
Metadata about the fields defined on the model, mapping of field names to [FieldInfo][pydantic.fields.FieldInfo].
This replaces Model.__fields__ from Pydantic V1.
- class pylinac.ct.CTP528Result(*, start_angle_radians: float, mtf_lp_mm: dict, roi_settings: dict)[source]#
Bases:
BaseModelThis class should not be called directly. It is returned by the
results_data()method. It is a dataclass under the hood and thus comes with all the dunder magic.Use the following attributes as normal class attributes.
Create a new model by parsing and validating input data from keyword arguments.
Raises [ValidationError][pydantic_core.ValidationError] if the input data cannot be validated to form a valid model.
self is explicitly positional-only to allow self as a field name.
- start_angle_radians: float#
- mtf_lp_mm: dict#
- roi_settings: dict#
- model_computed_fields: ClassVar[dict[str, ComputedFieldInfo]] = {}#
A dictionary of computed field names and their corresponding ComputedFieldInfo objects.
- model_config: ClassVar[ConfigDict] = {}#
Configuration for the model, should be a dictionary conforming to [ConfigDict][pydantic.config.ConfigDict].
- model_fields: ClassVar[dict[str, FieldInfo]] = {'mtf_lp_mm': FieldInfo(annotation=dict, required=True), 'roi_settings': FieldInfo(annotation=dict, required=True), 'start_angle_radians': FieldInfo(annotation=float, required=True)}#
Metadata about the fields defined on the model, mapping of field names to [FieldInfo][pydantic.fields.FieldInfo].
This replaces Model.__fields__ from Pydantic V1.
- class pylinac.ct.CTP515Result(*, cnr_threshold: float, num_rois_seen: int, roi_settings: dict, roi_results: dict)[source]#
Bases:
BaseModelThis class should not be called directly. It is returned by the
results_data()method. It is a dataclass under the hood and thus comes with all the dunder magic.Use the following attributes as normal class attributes.
Create a new model by parsing and validating input data from keyword arguments.
Raises [ValidationError][pydantic_core.ValidationError] if the input data cannot be validated to form a valid model.
self is explicitly positional-only to allow self as a field name.
- cnr_threshold: float#
- num_rois_seen: int#
- roi_settings: dict#
- roi_results: dict#
- model_computed_fields: ClassVar[dict[str, ComputedFieldInfo]] = {}#
A dictionary of computed field names and their corresponding ComputedFieldInfo objects.
- model_config: ClassVar[ConfigDict] = {}#
Configuration for the model, should be a dictionary conforming to [ConfigDict][pydantic.config.ConfigDict].
- model_fields: ClassVar[dict[str, FieldInfo]] = {'cnr_threshold': FieldInfo(annotation=float, required=True), 'num_rois_seen': FieldInfo(annotation=int, required=True), 'roi_results': FieldInfo(annotation=dict, required=True), 'roi_settings': FieldInfo(annotation=dict, required=True)}#
Metadata about the fields defined on the model, mapping of field names to [FieldInfo][pydantic.fields.FieldInfo].
This replaces Model.__fields__ from Pydantic V1.
- class pylinac.ct.CTP486Result(*, uniformity_index: float, integral_non_uniformity: float, nps_avg_power: float, nps_max_freq: float, passed: bool, rois: dict)[source]#
Bases:
BaseModelThis class should not be called directly. It is returned by the
results_data()method. It is a dataclass under the hood and thus comes with all the dunder magic.Use the following attributes as normal class attributes.
Create a new model by parsing and validating input data from keyword arguments.
Raises [ValidationError][pydantic_core.ValidationError] if the input data cannot be validated to form a valid model.
self is explicitly positional-only to allow self as a field name.
- uniformity_index: float#
- integral_non_uniformity: float#
- passed: bool#
- rois: dict#
- model_computed_fields: ClassVar[dict[str, ComputedFieldInfo]] = {}#
A dictionary of computed field names and their corresponding ComputedFieldInfo objects.
- model_config: ClassVar[ConfigDict] = {}#
Configuration for the model, should be a dictionary conforming to [ConfigDict][pydantic.config.ConfigDict].
- model_fields: ClassVar[dict[str, FieldInfo]] = {'integral_non_uniformity': FieldInfo(annotation=float, required=True), 'nps_avg_power': FieldInfo(annotation=float, required=True), 'nps_max_freq': FieldInfo(annotation=float, required=True), 'passed': FieldInfo(annotation=bool, required=True), 'rois': FieldInfo(annotation=dict, required=True), 'uniformity_index': FieldInfo(annotation=float, required=True)}#
Metadata about the fields defined on the model, mapping of field names to [FieldInfo][pydantic.fields.FieldInfo].
This replaces Model.__fields__ from Pydantic V1.
- class pylinac.ct.ROIResult(*, name: str, value: float, stdev: float, difference: float | None, nominal_value: float | None, passed: bool | None)[source]#
Bases:
BaseModelThis class should not be called directly. It is returned by the
results_data()method. It is a dataclass under the hood and thus comes with all the dunder magic.Use the following attributes as normal class attributes.
Create a new model by parsing and validating input data from keyword arguments.
Raises [ValidationError][pydantic_core.ValidationError] if the input data cannot be validated to form a valid model.
self is explicitly positional-only to allow self as a field name.
- name: str#
- value: float#
- stdev: float#
- difference: float | None#
- nominal_value: float | None#
- passed: bool | None#
- model_computed_fields: ClassVar[dict[str, ComputedFieldInfo]] = {}#
A dictionary of computed field names and their corresponding ComputedFieldInfo objects.
- model_config: ClassVar[ConfigDict] = {}#
Configuration for the model, should be a dictionary conforming to [ConfigDict][pydantic.config.ConfigDict].
- model_fields: ClassVar[dict[str, FieldInfo]] = {'difference': FieldInfo(annotation=Union[float, NoneType], required=True), 'name': FieldInfo(annotation=str, required=True), 'nominal_value': FieldInfo(annotation=Union[float, NoneType], required=True), 'passed': FieldInfo(annotation=Union[bool, NoneType], required=True), 'stdev': FieldInfo(annotation=float, required=True), 'value': FieldInfo(annotation=float, required=True)}#
Metadata about the fields defined on the model, mapping of field names to [FieldInfo][pydantic.fields.FieldInfo].
This replaces Model.__fields__ from Pydantic V1.
Module classes (CTP404, etc)#
- class pylinac.ct.Slice(catphan, slice_num: int | None = None, combine: bool = True, combine_method: str = 'mean', num_slices: int = 0, clear_borders: bool = True, original_image: ImageLike | None = None)[source]#
Bases:
objectBase class for analyzing specific slices of a CBCT dicom set.
Parameters#
- catphan
CatPhanBaseinstance. The catphan instance.
- slice_numint
The slice number of the DICOM array desired. If None, will use the
slice_numproperty of subclass.- combinebool
If True, combines the slices +/-
num_slicesaround the slice of interest to improve signal/noise.- combine_method{‘mean’, ‘max’}
How to combine the slices if
combineis True.- num_slicesint
The number of slices on either side of the nominal slice to combine to improve signal/noise; only applicable if
combineis True.- clear_bordersbool
If True, clears the borders of the image to remove any ROIs that may be present.
- original_image
Imageor None The array of the slice. This is a bolt-on parameter for optimization. Leaving as None is fine, but can increase analysis speed if 1) this image is passed and 2) there is no combination of slices happening, which is most of the time.
- property phantom_roi: RegionProperties#
Get the Scikit-Image ROI of the phantom
The image is analyzed to see if: 1) the CatPhan is even in the image (if there were any ROIs detected) 2) an ROI is within the size criteria of the catphan 3) the ROI area that is filled compared to the bounding box area is close to that of a circle
- catphan
- class pylinac.ct.CatPhanModule(catphan, tolerance: float | None = None, offset: int = 0, clear_borders: bool = True)[source]#
Bases:
SliceBase class for a CTP module.
Parameters#
- catphan
CatPhanBaseinstance. The catphan instance.
- slice_numint
The slice number of the DICOM array desired. If None, will use the
slice_numproperty of subclass.- combinebool
If True, combines the slices +/-
num_slicesaround the slice of interest to improve signal/noise.- combine_method{‘mean’, ‘max’}
How to combine the slices if
combineis True.- num_slicesint
The number of slices on either side of the nominal slice to combine to improve signal/noise; only applicable if
combineis True.- clear_bordersbool
If True, clears the borders of the image to remove any ROIs that may be present.
- original_image
Imageor None The array of the slice. This is a bolt-on parameter for optimization. Leaving as None is fine, but can increase analysis speed if 1) this image is passed and 2) there is no combination of slices happening, which is most of the time.
- roi_dist_mm#
alias of
float
- roi_radius_mm#
alias of
float
- catphan
- class pylinac.ct.CTP404CP503(catphan, offset: int, hu_tolerance: float, thickness_tolerance: float, scaling_tolerance: float, clear_borders: bool = True, thickness_slice_straddle: str | int = 'auto', expected_hu_values: dict[str, float | int] | None = None)[source]#
Bases:
CTP404CP504Alias for namespace consistency
Parameters#
catphan : ~pylinac.cbct.CatPhanBase instance. offset : int hu_tolerance : float thickness_tolerance : float scaling_tolerance : float clear_borders : bool
- class pylinac.ct.CTP404CP504(catphan, offset: int, hu_tolerance: float, thickness_tolerance: float, scaling_tolerance: float, clear_borders: bool = True, thickness_slice_straddle: str | int = 'auto', expected_hu_values: dict[str, float | int] | None = None)[source]#
Bases:
CatPhanModuleClass for analysis of the HU linearity, geometry, and slice thickness regions of the CTP404.
Parameters#
catphan : ~pylinac.cbct.CatPhanBase instance. offset : int hu_tolerance : float thickness_tolerance : float scaling_tolerance : float clear_borders : bool
- preprocess(catphan) None[source]#
A preprocessing step before analyzing the CTP module.
Parameters#
catphan : ~pylinac.cbct.CatPhanBase instance.
- property lcv: float#
The low-contrast visibility
- plot_linearity(axis: Axes | None = None, plot_delta: bool = True) tuple[source]#
Plot the HU linearity values to an axis.
Parameters#
- axisNone, matplotlib.Axes
The axis to plot the values on. If None, will create a new figure.
- plot_deltabool
Whether to plot the actual measured HU values (False), or the difference from nominal (True).
- property passed_hu: bool#
Boolean specifying whether all the ROIs passed within tolerance.
- property passed_thickness: bool#
Whether the slice thickness was within tolerance from nominal.
- property meas_slice_thickness: float#
The average slice thickness for the 4 wire measurements in mm.
- property passed_geometry: bool#
Returns whether all the line lengths were within tolerance.
- class pylinac.ct.CTP404CP600(catphan, offset: int, hu_tolerance: float, thickness_tolerance: float, scaling_tolerance: float, clear_borders: bool = True, thickness_slice_straddle: str | int = 'auto', expected_hu_values: dict[str, float | int] | None = None)[source]#
Bases:
CTP404CP504Parameters#
catphan : ~pylinac.cbct.CatPhanBase instance. offset : int hu_tolerance : float thickness_tolerance : float scaling_tolerance : float clear_borders : bool
- class pylinac.ct.CTP404CP604(catphan, offset: int, hu_tolerance: float, thickness_tolerance: float, scaling_tolerance: float, clear_borders: bool = True, thickness_slice_straddle: str | int = 'auto', expected_hu_values: dict[str, float | int] | None = None)[source]#
Bases:
CTP404CP504Parameters#
catphan : ~pylinac.cbct.CatPhanBase instance. offset : int hu_tolerance : float thickness_tolerance : float scaling_tolerance : float clear_borders : bool
- class pylinac.ct.CTP528CP503(catphan, tolerance: float | None = None, offset: int = 0, clear_borders: bool = True)[source]#
Bases:
CTP528CP504Parameters#
- catphan
CatPhanBaseinstance. The catphan instance.
- slice_numint
The slice number of the DICOM array desired. If None, will use the
slice_numproperty of subclass.- combinebool
If True, combines the slices +/-
num_slicesaround the slice of interest to improve signal/noise.- combine_method{‘mean’, ‘max’}
How to combine the slices if
combineis True.- num_slicesint
The number of slices on either side of the nominal slice to combine to improve signal/noise; only applicable if
combineis True.- clear_bordersbool
If True, clears the borders of the image to remove any ROIs that may be present.
- original_image
Imageor None The array of the slice. This is a bolt-on parameter for optimization. Leaving as None is fine, but can increase analysis speed if 1) this image is passed and 2) there is no combination of slices happening, which is most of the time.
- catphan
- class pylinac.ct.CTP528CP504(catphan, tolerance: float | None = None, offset: int = 0, clear_borders: bool = True)[source]#
Bases:
CatPhanModuleClass for analysis of the Spatial Resolution slice of the CBCT dicom data set.
A collapsed circle profile is taken of the line-pair region. This profile is search for peaks and valleys. The MTF is calculated from those peaks & valleys.
Attributes#
- radius2linepairs_mmfloat
The radius in mm to the line pairs.
Parameters#
- catphan
CatPhanBaseinstance. The catphan instance.
- slice_numint
The slice number of the DICOM array desired. If None, will use the
slice_numproperty of subclass.- combinebool
If True, combines the slices +/-
num_slicesaround the slice of interest to improve signal/noise.- combine_method{‘mean’, ‘max’}
How to combine the slices if
combineis True.- num_slicesint
The number of slices on either side of the nominal slice to combine to improve signal/noise; only applicable if
combineis True.- clear_bordersbool
If True, clears the borders of the image to remove any ROIs that may be present.
- original_image
Imageor None The array of the slice. This is a bolt-on parameter for optimization. Leaving as None is fine, but can increase analysis speed if 1) this image is passed and 2) there is no combination of slices happening, which is most of the time.
- property mtf: MTF#
The Relative MTF of the line pairs, normalized to the first region.
Returns#
dict
- property radius2linepairs: float#
Radius from the phantom center to the line-pair region, corrected for pixel spacing.
- property circle_profile: CollapsedCircleProfile#
Calculate the median profile of the Line Pair region.
Returns#
pylinac.core.profile.CollapsedCircleProfile: A 1D profile of the Line Pair region.
- class pylinac.ct.CTP528CP600(catphan, tolerance: float | None = None, offset: int = 0, clear_borders: bool = True)[source]#
Bases:
CTP528CP504Parameters#
- catphan
CatPhanBaseinstance. The catphan instance.
- slice_numint
The slice number of the DICOM array desired. If None, will use the
slice_numproperty of subclass.- combinebool
If True, combines the slices +/-
num_slicesaround the slice of interest to improve signal/noise.- combine_method{‘mean’, ‘max’}
How to combine the slices if
combineis True.- num_slicesint
The number of slices on either side of the nominal slice to combine to improve signal/noise; only applicable if
combineis True.- clear_bordersbool
If True, clears the borders of the image to remove any ROIs that may be present.
- original_image
Imageor None The array of the slice. This is a bolt-on parameter for optimization. Leaving as None is fine, but can increase analysis speed if 1) this image is passed and 2) there is no combination of slices happening, which is most of the time.
- catphan
- class pylinac.ct.CTP528CP604(catphan, tolerance: float | None = None, offset: int = 0, clear_borders: bool = True)[source]#
Bases:
CTP528CP504Alias for namespace consistency.
Parameters#
- catphan
CatPhanBaseinstance. The catphan instance.
- slice_numint
The slice number of the DICOM array desired. If None, will use the
slice_numproperty of subclass.- combinebool
If True, combines the slices +/-
num_slicesaround the slice of interest to improve signal/noise.- combine_method{‘mean’, ‘max’}
How to combine the slices if
combineis True.- num_slicesint
The number of slices on either side of the nominal slice to combine to improve signal/noise; only applicable if
combineis True.- clear_bordersbool
If True, clears the borders of the image to remove any ROIs that may be present.
- original_image
Imageor None The array of the slice. This is a bolt-on parameter for optimization. Leaving as None is fine, but can increase analysis speed if 1) this image is passed and 2) there is no combination of slices happening, which is most of the time.
- catphan
- class pylinac.ct.CTP515(catphan, tolerance: float, cnr_threshold: float, offset: int, contrast_method: str, visibility_threshold: float, clear_borders: bool = True)[source]#
Bases:
CatPhanModuleClass for analysis of the low contrast slice of the CTP module. Low contrast is measured by obtaining the average pixel value of the contrast ROIs and comparing that value to the average background value. To obtain a more “human” detection level, the contrast (which is largely the same across different-sized ROIs) is multiplied by the diameter. This value is compared to the contrast threshold to decide if it can be “seen”.
Parameters#
- catphan
CatPhanBaseinstance. The catphan instance.
- slice_numint
The slice number of the DICOM array desired. If None, will use the
slice_numproperty of subclass.- combinebool
If True, combines the slices +/-
num_slicesaround the slice of interest to improve signal/noise.- combine_method{‘mean’, ‘max’}
How to combine the slices if
combineis True.- num_slicesint
The number of slices on either side of the nominal slice to combine to improve signal/noise; only applicable if
combineis True.- clear_bordersbool
If True, clears the borders of the image to remove any ROIs that may be present.
- original_image
Imageor None The array of the slice. This is a bolt-on parameter for optimization. Leaving as None is fine, but can increase analysis speed if 1) this image is passed and 2) there is no combination of slices happening, which is most of the time.
- property rois_visible: int#
The number of ROIs “visible”.
- property window_min: float#
Lower bound of CT window/leveling to show on the plotted image. Improves apparent contrast.
- property window_max: float#
Upper bound of CT window/leveling to show on the plotted image. Improves apparent contrast
- catphan
- class pylinac.ct.CTP486(catphan, tolerance: float | None = None, offset: int = 0, clear_borders: bool = True)[source]#
Bases:
CatPhanModuleClass for analysis of the Uniformity slice of the CTP module. Measures 5 ROIs around the slice that should all be close to the same value.
Parameters#
- catphan
CatPhanBaseinstance. The catphan instance.
- slice_numint
The slice number of the DICOM array desired. If None, will use the
slice_numproperty of subclass.- combinebool
If True, combines the slices +/-
num_slicesaround the slice of interest to improve signal/noise.- combine_method{‘mean’, ‘max’}
How to combine the slices if
combineis True.- num_slicesint
The number of slices on either side of the nominal slice to combine to improve signal/noise; only applicable if
combineis True.- clear_bordersbool
If True, clears the borders of the image to remove any ROIs that may be present.
- original_image
Imageor None The array of the slice. This is a bolt-on parameter for optimization. Leaving as None is fine, but can increase analysis speed if 1) this image is passed and 2) there is no combination of slices happening, which is most of the time.
- plot_profiles(axis: Axes | None = None) None[source]#
Plot the horizontal and vertical profiles of the Uniformity slice.
Parameters#
- axisNone, matplotlib.Axes
The axis to plot on; if None, will create a new figure.
- property overall_passed: bool#
Boolean specifying whether all the ROIs passed within tolerance.
- property uniformity_index: float#
The Uniformity Index. Elstrom et al equation 2. https://www.tandfonline.com/doi/pdf/10.3109/0284186X.2011.590525
- property integral_non_uniformity: float#
The Integral Non-Uniformity. Elstrom et al equation 1. https://www.tandfonline.com/doi/pdf/10.3109/0284186X.2011.590525
- property power_spectrum_2d: ndarray#
The power spectrum of the uniformity ROI.
- property power_spectrum_1d: ndarray#
The 1D power spectrum of the uniformity ROI.
- property avg_noise_power: float#
The average noise power of the uniformity ROI.
- property max_noise_power_frequency: float#
The frequency of the maximum noise power. 0 means no pattern.
- catphan
ROI Objects#
- class pylinac.ct.HUDiskROI(array: ndarray | ArrayImage, angle: float, roi_radius: float, dist_from_center: float, phantom_center: tuple | Point, nominal_value: float | None = None, tolerance: float | None = None, background_mean: float | None = None, background_std: float | None = None)[source]#
Bases:
DiskROIAn HU ROI object. Represents a circular area measuring either HU sample (Air, Poly, …) or HU uniformity (bottom, left, …).
Parameters#
- nominal_value
The nominal pixel value of the HU ROI.
- tolerance
The roi pixel value tolerance.
- property value_diff: float#
The difference in HU between measured and nominal.
- property passed: bool#
Boolean specifying if ROI pixel value was within tolerance of the nominal value.
- property plot_color: str#
Return one of two colors depending on if ROI passed.
- class pylinac.ct.ThicknessROI(array, width, height, angle, dist_from_center, phantom_center)[source]#
Bases:
RectangleROIA rectangular ROI that measures the angled wire rod in the HU linearity slice which determines slice thickness.
Parameters#
- widthnumber
Width of the rectangle. Must be positive
- heightnumber
Height of the rectangle. Must be positive.
- centerPoint, iterable, optional
Center point of rectangle.
- as_intbool
If False (default), inputs are left as-is. If True, all inputs are converted to integers.
- property long_profile: FWXMProfile#
The profile along the axis perpendicular to ramped wire.
- property wire_fwhm: float#
The FWHM of the wire in pixels.
- property plot_color: str#
The plot color.
- class pylinac.ct.GeometricLine(geo_roi1: Point, geo_roi2: Point, mm_per_pixel: float, tolerance: int | float)[source]#
Bases:
LineRepresents a line connecting two nodes/ROIs on the Geometry Slice.
Attributes#
- nominal_length_mmint, float
The nominal distance between the geometric nodes, in mm.
Parameters#
- geo_roi1GEO_ROI
One of two ROIs representing one end of the line.
- geo_roi2GEO_ROI
The other ROI which is the other end of the line.
- mm_per_pixelfloat
The mm/pixel value.
- toleranceint, float
The tolerance of the geometric line, in mm.
- property passed: bool#
Whether the line passed tolerance.
- property pass_fail_color: str#
Plot color for the line, based on pass/fail status.
- property length_mm: float#
Return the length of the line in mm.
Helper Functions#
- pylinac.ct.combine_surrounding_slices(dicomstack: DicomImageStack, nominal_slice_num: int, slices_plusminus: int = 1, mode: str = 'mean') array[source]#
Return an array that is the combination of a given slice and a number of slices surrounding it.
Parameters#
- dicomstack~pylinac.core.image.DicomImageStack
The CBCT DICOM stack.
- nominal_slice_numint
The slice of interest (along 3rd dim).
- slices_plusminus: int
How many slices plus and minus to combine (also along 3rd dim).
- mode{‘mean’, ‘median’, ‘max}
Specifies the method of combination.
Returns#
- combined_arraynumpy.array
The combined array of the DICOM stack slices.